Allocate Samples to Batches Using Specified Method
allocate_samples.RdThis function allocates samples to batches based on the specified method, which can be random allocation, best random allocation, or simulated annealing. It supports optional blocking and handles various types of covariates.
Usage
allocate_samples(
data,
id_column = "sample_id",
method = "simulated_annealing",
covariates,
blocking_variable = NA,
batch_size,
iterations = 1000,
temperature = 1,
cooling_rate = 0.975,
seed = 123,
plot_convergence = TRUE,
collapse_rare_factors = FALSE,
balance_metric = "harmonic_mean",
balance_type = "p_value"
)Arguments
- data
A
data.framecontaining the dataset to be processed.- id_column
A string specifying the column name in
datathat contains the sample IDs. The default is "sample_id".- method
A string specifying the allocation method to be used. Valid options are "random", "best_random", and "simulated_annealing". The default is "simulated_annealing".
- covariates
A character vector listing the names of the covariate columns in
datathat should be considered during allocation.- blocking_variable
An optional string specifying the name of the column to be used as a blocking variable. If not provided or
NA, blocking is not applied.- batch_size
An integer specifying the size of each batch.
- iterations
An integer specifying the number of iterations to run for the "best_random" or "simulated_annealing" methods. The default is 1000.
- temperature
A numeric specifying the initial temperature for the simulated annealing method. The default is 1.
- cooling_rate
A numeric specifying the cooling rate for the simulated annealing method. The default is 0.975.
- seed
An integer used for setting the seed to ensure reproducibility. The default is 123.
- plot_convergence
= TRUE A logical indicating whether to plot the convergence of the optimisation process, only relevant if the method specified is "simulated_annealing". The default is
TRUE.- collapse_rare_factors
= FALSE A logical indicating whether to collapse rare factor levels in the covariates. The default is
FALSE. Factor levels that are present with counts either less than 5 or in 0.05*batch_size are collapsed into a single level "other". Please note that the layout returned will contain these collapsed factor levels, rather than your original input data.- balance_metric
A character string specifying how to combine multiple covariate metrics. Valid options are:
"harmonic_mean"(default): Non-compensatory combination. Ensures all covariates must be balanced - no single well-balanced covariate can compensate for a poorly balanced one. More sensitive to the worst-balanced covariate."product": Product of all metrics. Very sensitive to any poorly balanced covariate. Can become numerically small with many covariates.
- balance_type
A character string specifying what metric to calculate for each covariate. Valid options are:
"p_value"(default): Uses p-values from statistical tests (Kruskal-Wallis for continuous covariates, Fisher's exact test for categorical covariates). Tests whether covariate distributions differ significantly between batches. Higher p-values indicate better balance."effect_size": Uses effect sizes measuring the magnitude of association between covariates and batch assignment. Uses Cramér's V for categorical covariates and eta (from ANOVA) for continuous covariates. Effect sizes range from 0 (no association, perfect balance) to 1 (maximum association). Note: effect sizes are internally transformed as (1 - effect_size) so that higher values indicate better balance, consistent with p-values.
Value
A named list with elements:
- layout
Data frame with original data plus
batch_allocationcolumn- results
Data frame with covariate balance metrics (
covariate,p_value,effect_size)- optimisation_data
(Only for iterative methods) Data frame with
iteration,temperature,objective_value
Details
The function first checks the validity of the specified method and parameters, then preprocesses the data according to the specified covariates and blocking variable. It applies the specified allocation method to assign samples to batches, aiming to balance the distribution of covariates and, if applicable, blocking levels across batches.
Examples
# Allocate samples using simulated annealing without blocking
my_data = simulate_data(n_samples = 100)
allocated_data <- allocate_samples(data = my_data,
id_column = "sample_id",
method = "simulated_annealing",
covariates = c("age_at_baseline", "bmi_at_baseline", "sex"),
batch_size = 13)
#> No blocking variable specified.
#> Covariate: age_at_baseline - continuous
#> Covariate: bmi_at_baseline - continuous
#> Covariate: sex - categorical
#> Number of samples: 100
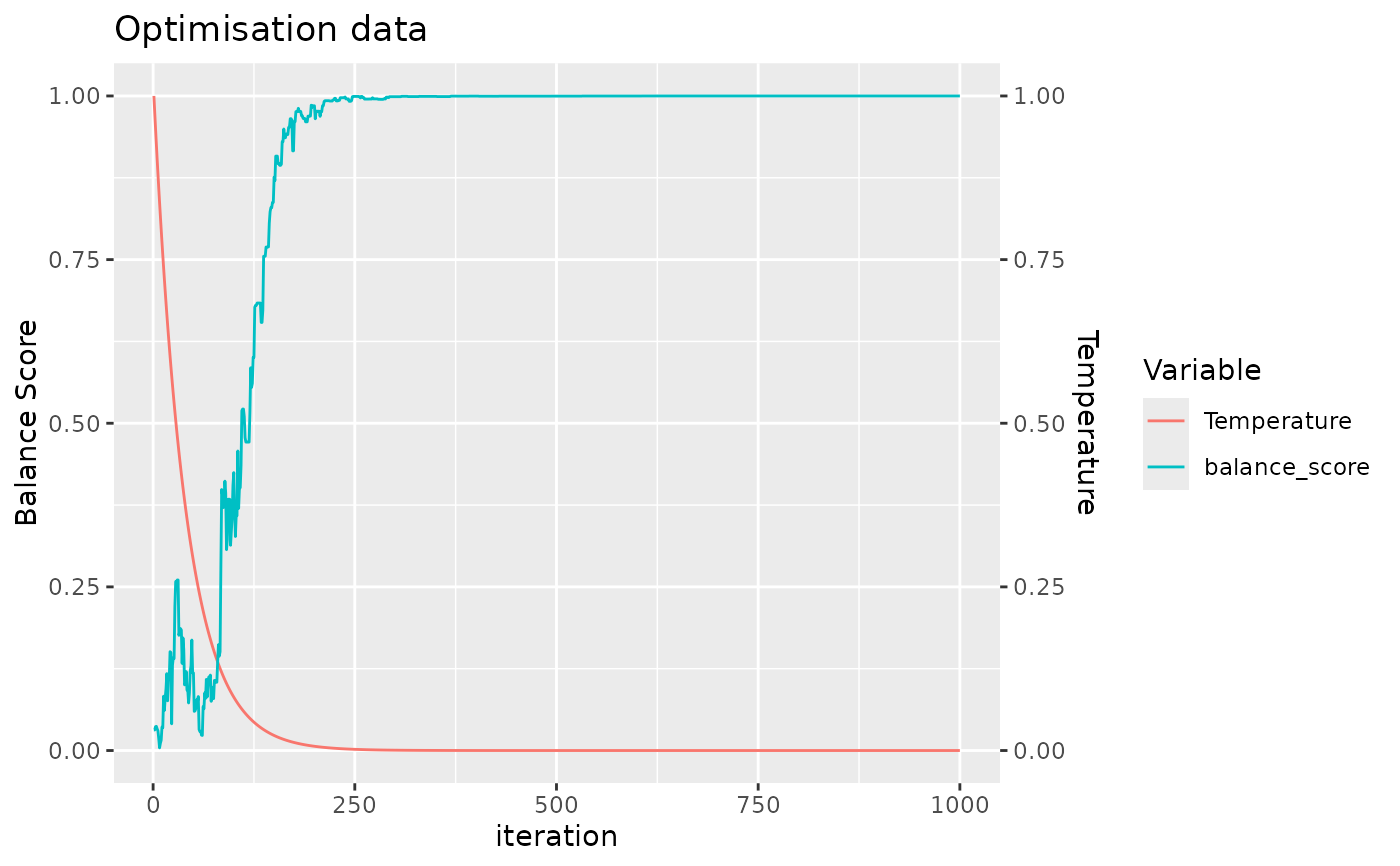 #> Balance Score of final layout: 1
#> Joining with `by = join_by(age_at_baseline, bmi_at_baseline, sex, sample_id)`
#> Balance Score of final layout: 1
#> Joining with `by = join_by(age_at_baseline, bmi_at_baseline, sex, sample_id)`